
PubReading
343 episodes — Page 7 of 7
S1 Ep 38PubReading [43] - Click and Cut: a click chemistry approach to developing oxidative DNA damaging agents - N. McStay, A. Kellett et al.
Metallodrugs provide important first-line treatment against various forms of human cancer. To overcome chemotherapeutic resistance and widen treatment possibilities, new agents with improved or alternative modes of action are highly sought after. Here, we present a click chemistry strategy for developing DNA damaging metallodrugs. The approach in- volves the development of a series of polyamine ligands where three primary, secondary or tertiary alkyne-amines were selected and ‘clicked’ using the copper-catalysed azide-alkyne cycloaddition reac- tion to a 1,3,5-azide mesitylene core to produce a family of compounds we call the ‘Tri-Click’ (TC) se- ries. From the isolated library, one dominant ligand (TC1) emerged as a high-affinity copper(II) binding agent with potent DNA recognition and damaging properties. Using a range of in vitro biophysical and molecular techniques––including free radical scavengers, spin trapping antioxidants and base excision repair (BER) enzymes–the oxidative DNA damaging mechanism of copper-bound TC1 was elucidated. This activity was then compared to intracellular results obtained from peripheral blood mononuclear cells exposed to Cu(II)–TC1 where use of BER enzymes and fluorescently modified dNTPs enabled the characterisation and quantification of genomic DNA lesions produced by the complex. The approach can serve as a new avenue for the design of DNA damaging agents with unique activity profiles. - doi.org/10.1093/nar/gkab817 - 2021
S1 Ep 37PubReading [42] - Circulating tumoral DNA: Preanalytical validation and quality control in a diagnostic laboratory - S. Nikolaev, T. Nouspikel et al.
We present the results of our technical validation process in establishing the analysis of circulating tumor DNA (ctDNA) as a diagnostic tool. Like most cells in our body, tumor cells shed DNA in the blood flow. Analysis of ctDNA mutational content can provide invaluable information on the genetic makeup of a tumor, and assist oncologists in deciding on therapy, or in following residual disease. However, low absolute amounts of circulating DNA and low tumor fraction constitute formidable analytical challenges. A key step is to avoid contamination with genomic DNA from cell lysis. Several brands of specialized blood collection tubes are available to prevent leukocyte lysis. We show that they are not equally efficient, depending on storage temperature and time before plasma preparation. We report our analysis of preanalytical factors pertaining to ctDNA analysis (tubes, transportation time, temperature) and our conclusions in terms of instructions to prescribing physicians. We also stress the importance of proper DNA quality control and compare several methods, including a differential amplicon length PCR technique which allows determination of multiple QC parameters from minimal amounts of DNA. Altogether, these data provide useful practical information to diagnostic laboratories wishing to implement the assay of ctDNA in clinical practice. - doi.org/10.1016/j.ab.2017.11.004 - 2018
S1 Ep 36PubReading [41] - Apoptosis: A Target for Anticancer Therapy - C. M. Pfeffer and A. T. K. Singh
Apoptosis, the cell’s natural mechanism for death, is a promising target for anticancer therapy. Both the intrinsic and extrinsic pathways use caspases to carry out apoptosis through the cleavage of hundreds of proteins. In cancer, the apoptotic pathway is typically inhibited through a wide variety of means including overexpression of antiapoptotic proteins and under-expression of proapoptotic proteins. Many of these changes cause intrinsic resistance to the most common anticancer therapy, chemotherapy. Promising new anticancer therapies are plant-derived compounds that exhibit anticancer activity through activating the apoptotic pathway. doi:10.3390/ijms19020448 - 2018
S1 Ep 35PubReading [40] - Phosphoproteome profiling uncovers a key role for CDKs in TNF signaling - M. Tanzer, M. Mann et al
Tumor necrosis factor (TNF) is one of the few cytokines successfully targeted by therapies against inflammatory diseases. However, blocking this well studied and pleiotropic ligand can cause dramatic side-effects. Here, we reason that a systems-level proteomic analysis of TNF signaling could dissect its diverse functions and offer a base for developing more targeted therapies. Therefore, we combine phosphoproteomics time course experiments with sub-cellular localization and kinase inhibitor analysis to identify functional modules of protein phosphorylation. The majority of regulated phosphorylation events can be assigned to an upstream kinase by inhibiting master kinases. Spatial proteomics reveals phosphorylation- dependent translocations of hundreds of proteins upon TNF stimulation. Phosphoproteome analysis of TNF-induced apoptosis and necroptosis uncovers a key role for transcriptional cyclin-dependent kinase activity to promote cytokine production and prevent excessive cell death downstream of the TNF signaling receptor. This resource of TNF-induced pathways and sites can be explored at http://tnfviewer.biochem.mpg.de/. - doi.org/10.1038/s41467-021-26289-6 - 2021
S1 Ep 34PubReading [39] - Crystal structure of a membrane-bound metalloenzyme that catalyses the biological oxidation of methane - R. L. Lieberman and A. C. Rosenzweig
Particulate methane monooxygenase (pMMO) is an integral membrane metalloenzyme that catalyses the conversion of methane to methanol. Knowledge of how pMMO performs this extremely challenging chemistry may have an impact on the use of methane as an alternative energy source by facilitating the development of new synthetic catalysts. We have determined the structure of pMMO from the methanotroph Methylococcus capsulatus (Bath) to a resolution of 2.8A ̊. The enzyme is a trimer with an a3b3g3 polypeptide arrangement. Two metal centres, modelled as mononuclear copper and dinuclear copper, are located in soluble regions of each pmoB subunit, which resembles cytochrome c oxidase subunit II. A third metal centre, occupied by zinc in the crystal, is located within the membrane. The structure provides new insight into the molecular details of biological methane oxidation. - doi.org/10.1038/nature03311 - 2005
S1 Ep 33PubReading [38] - PeptiCHIP: A Microfluidic Platform for Tumor Antigen Landscape Identification - S. Feola, V. Cerullo et al.
Identification of HLA class I ligands from the tumor surface (ligandome or immunopeptidome) is essential for designing T-cell mediated cancer therapeutic approaches. However, the sensitivity of the process for isolating MHC-I restricted tumor-specific peptides has been the major limiting factor for reliable tumor antigen characterization, making clear the need for technical improvement. Here, we describe our work from the fabrication and development of a microfluidic-based chip (PeptiCHIP) and its use to identify and characterize tumor-specific ligands on clinically relevant human samples. Specifically, we assessed the potential of immobilizing a pan-HLA antibody on solid surfaces via well-characterized streptavidin−biotin chemistry, overcoming the limitations of the cross-linking chemistry used to prepare the affinity matrix with the desired antibodies in the immunopeptidomics workflow. Furthermore, to address the restrictions related to the handling and the limited availability of tumor samples, we further developed the concept toward the implementation of a microfluidic through-flow system. Thus, the biotinylated pan-HLA antibody was immobilized on streptavidin-functionalized surfaces, and immune-affinity purification (IP) was carried out on customized microfluidic pillar arrays made of thiol−ene polymer. Compared to the standard methods reported in the field, our methodology reduces the amount of antibody and the time required for peptide isolation. In this work, we carefully examined the specificity and robustness of our customized technology for immunopeptidomics workflows. We tested this platform by immunopurifying HLA-I complexes from 1 × 106 cells both in a widely studied B-cell line and in patients-derived ex vivo cell cultures, instead of 5 × 108 cells as required in the current technology. After the final elution in mild acid, HLA-I-presented peptides were identified by tandem mass spectrometry and further investigated by in vitro methods. These results highlight the potential to exploit microfluidics-based strategies in immunopeptidomics platforms and in personalized immunopeptidome analysis from cells isolated from individual tumor biopsies to design tailored cancer therapeutic vaccines. Moreover, the possibility to integrate multiple identical units on a single chip further improves the throughput and multiplexing of these assays with a view to clinical needs. - doi.org/10.1021/acsnano.1c04371 - 2021
S1 Ep 32PubReading [37] - High resolution single particle cryo-electron microscopy using beam-image shift - A. Cheng, B. Carragher et al.
Automated data acquisition is used widely for single-particle reconstruction of three-dimensional (3D) volumes of biological complexes preserved in vitreous ice and imaged in a transmission electron microscope. Automation has become integral to this method because of the very large number of particle images required in order to overcome the typically low signal-to-noise ratio of these images. For optimal efficiency, automated data acquisition software packages typically employ some beam-image shift targeting as this method is both fast and accurate ( ± 0.1 μm). In contrast, using only stage movement, relocation to a targeted area under low-dose conditions can only be achieved in combination with multiple iterations or long relaxation times, both reducing efficiency. Nevertheless it is well known that applying beam- image shift induces beam-tilt and with it a potential structure phase error with a phase error π/4 the highest acceptable value. This theory has been used as an argument against beam-image shift for high resolution data collection. Nevertheless, in practice many small beam-image shift datasets have resulted in 3D reconstructions beyond the π/4 phase error limit. To address this apparent contradiction, we performed cryo-EM single-particle reconstructions on a T20S proteasome sample using applied beam-image shifts corresponding to beam tilts from 0 to 10 mrad. To evaluate the results we compared the FSC values, and examined the water density peaks in the 3D map. We conclude that the phase error does not limit the validity of the 3D reconstruction from single-particle averaging beyond the π/4 resolution limit. - doi.org/10.1016/j.jsb.2018.07.015 - 2018
S1 Ep 31PubReading [36] - APE1: A skilled nucleic acid surgeon - A. Whitaker and B. Freudenthal
Before a deleterious DNA lesion can be replaced with its undamaged counterpart, the lesion must first be removed from the genome. This process of removing and replacing DNA lesions is accomplished by the careful coordination of several protein factors during DNA repair. One such factor is the multifunctional enzyme human apurinic/apyrimidinic endonuclease 1 (APE1), known best for its DNA backbone cleavage activity at AP sites during base excision repair (BER). APE1 preforms AP site incision with surgical precision and skill, by sculpting the DNA to place the cleavage site in an optimal position for nucleophilic attack within its compact protein active site. APE1, however, has demonstrated broad surgical expertise, and applies its DNA cleavage activity to a wide variety of DNA and RNA substrates. Here, we discuss what is known and unknown about APE1 cleavage mechanisms, focusing on structural and mechanistic considerations. Importantly, disruptions in the biological functions associated with APE1 are linked to numerous human maladies, including cancer and neurodegenerative diseases. The continued elucidation of APE1 mechanisms is required for rational drug design towards novel and strategic ways to target its associated repair pathways. - doi.org/10.1016/j.dnarep.2018.08.012 - 2018
S1 Ep 30PubReading [35] - Oligonucleotide conjugated multi-functional adeno-associated viruses - D. Katrekar, P. Mali et al.
Recombinant adeno-associated viruses (AAVs) are among the most commonly used vehicles for in vivo gene delivery. However, their tropism is limited, and additionally their efficacy can be negatively affected by prevalence of neutralizing antibodies in sera. Methodologies to systematically engineer AAV capsid properties would thus be of great relevance. In this regard, we develop here multi-functional AAVs by engineering precision tethering of oligonucleotides onto the AAV surface, and thereby enabling a spectrum of nucleic-acid programmable functionalities. Towards this, we engineered genetically encoded incorporation of unnatural amino acids (UAA) bearing bio-orthogonal chemical handles onto capsid proteins. Via these we enabled site-specific coupling of oligonucleotides onto the AAV capsid surface using facile click chemistry. The resulting oligo-AAVs could be sequence specifically labeled, and also patterned in 2D using DNA array substrates. Additionally, we utilized these oligo conjugations to engineer viral shielding by lipid-based cloaks that efficaciously protected the AAV particles from neutralizing serum. We confirmed these ‘cloaked AAVs’ retained full functionality via their ability to transduce a range of cell types, and also enable robust delivery of CRISPR-Cas9 effectors. Taken together, we anticipate this programmable oligo-AAV system will have broad utility in synthetic biology and AAV engineering applications. - DOI:10.1038/s41598-018-21742-x - 2018
S1 Ep 29PubReading [34] - Replication protein A binds RNA and promotes R-Loop formation - O. Mazina, A. V. Mazin et al.
Replication protein A (RPA), a major eukaryotic ssDNA- binding protein, is essential for all metabolic processes that involve ssDNA, including DNA replication, repair, and damage signaling. To perform its functions, RPA binds ssDNA tightly. In contrast, it was presumed that RPA binds RNA weakly. However, recent data suggest that RPA may play a role in RNA metabolism. RPA stimulates RNA-templated DNA repair in vitro and associates in vivo with R-loops, the three-stranded structures consisting of an RNA-DNA hybrid and the displaced ssDNA strand. R-loops are common in the genomes of pro- and eukaryotes, including humans, and may play an important role in transcription-coupled homologous recombination and DNA replication restart. However, the mechanism of R-loop formation remains unknown. Here, we investigated the RNA-binding properties of human RPA and its possible role in R-loop formation. Using gel-retardation and RNA/ DNA competition assays, we found that RPA binds RNA with an unexpectedly high affinity (KD ’ 100 pM). Furthermore, RPA, by forming a complex with RNA, can promote R-loop formation with homologous dsDNA. In reconstitution experiments, we showed that human DNA polymerases can utilize RPA-generated R-loops for initiation of DNA synthesis, mimicking the process of replication restart in vivo. These results demonstrate that RPA binds RNA with high affinity, supporting the role of this protein in RNA metabolism and suggesting a mechanism of genome maintenance that depends on RPA- mediated DNA replication restart. - DOI 10.1074/jbc.RA120.013812 - 2020
S1 Ep 28PubReading [33] - Oxidative DNA Cleavage with Clip-Phenanthroline Triplex-Forming Oligonucleotide Hybrids - A. Panattoni, M. Hocek et al.
A systematic study of several new types of hybrids of Cu-chelated clamped phenanthroline artificial metallonuclease (AMN) with triplex-forming oligonucleotides (TFO) for sequence-specific cleavage of double-stranded DNA (dsDNA) is reported. The synthesis of these AMN–TFO hybrids is based on application of the alkyne–azide cycloaddition click reaction as the key step. The AMN was attached through different linkers at either the 5’- or 3’-ends or in the middle of the TFO stretch. The diverse hybrids efficiently formed triplexes with the target purine-rich sequence and their copper complexes were studied for their ability to cleave dsDNA in the presence of ascorbate as a reductant. In all cases, the influence of the nature and length of the AMN–TFO, time, conditions and amounts of ascorbate were studied, and optimum conjugates and a procedure that gave reasonably efficient (up to 34%) cleavage of the target sequence, while rendering an off-target dsDNA intact, were found. The footprint of cleavage on PAGE was identified only in one case, with low conversion; this means that cleavage does not proceed with single nucleotide precision. On the other hand, these AMN–TFO hybrids are useful for the selective degradation of target dsDNA sequences. Future improvements to this design may provide higher resolution and selectivity. - doi.org/10.1002/cbic.201900670. - 2019
S1 Ep 27PubReading [32] - Enhancing Protein Crystallization under a Magnetic Field - S. Young Ryu, H. Kyu Song et al.
High-quality crystals are essential to ensure high-resolution structural information. Protein crystals are controlled by many factors, such as pH, temperature, and the ion concentration of crystalline solutions. We previously reported the development of a device dedicated to protein crystallization. In the current study, we have further modified and improved our device. Exposure to external magnetic field leads to alignment of the crystal toward a preferred direction depending on the magnetization energy. Each material has different magnetic susceptibilities depending on the individual direction of their unit crystal cells. One of the strategies to acquire a large crystal entails controlling the nucleation rate. Furthermore, exposure of a crystal to a magnetic field may lead to new morphologies by affecting the crystal volume, shape, and quality. - doi:10.3390/cryst10090821 - 2020
S1 Ep 26PubReading [31] - TLR8 Is a Sensor of RNase T2 Degradation Products - W. Greulic, T. Carell, V. Hornung et al.
TLR8 is among the highest-expressed pattern-recognition receptors in the human myeloid compartment, yet its mode of action is poorly understood. TLR8 engages two distinct ligand binding sites to sense RNA degradation products, although it remains unclear how these ligands are formed in cellulo in the context of complex RNA molecule sensing. Here, we identified the lysosomal endoribonuclease RNase T2 as a non-redundant upstream component of TLR8- dependent RNA recognition. RNase T2 activity is required for rendering complex single-stranded, exogenous RNA molecules detectable for TLR8. This is due to RNase T2’s preferential cleavage of single-stranded RNA molecules between purine and uridine residues, which critically contributes to the supply of catabolic uridine and the generation of purine-2',3'-cyclophosphate-terminated oligoribonucleotides. Thus-generated molecules constitute agonistic ligands for the first and second binding pocket of TLR8. Together, these results establish the identity and origin of the RNA-derived molecular pattern sensed by TLR8. - doi.org/10.1016/j.cell.2019.11.001 - 2019
S1 Ep 25PubReading [30] - Cone-shaped HIV-1 capsids are transported through intact nuclear pores - V. Zila, M. Beck et al.
Human immunodeficiency virus (HIV-1) remains a major health threat. Viral capsid uncoating and nuclear import of the viral genome are critical for productive infection. The size of the HIV-1 capsid is generally believed to exceed the diameter of the nuclear pore complex (NPC), indicating that capsid uncoating has to occur prior to nuclear import. Here, we combined correlative light and electron microscopy with subtomogram averaging to capture the structural status of reverse transcription-competent HIV-1 complexes in infected T cells. We demonstrated that the diameter of the NPC in cellulo is sufficient for the import of apparently intact, cone-shaped capsids. Subsequent to nuclear import, we detected disrupted and empty capsid fragments, indicating that uncoating of the replication complex occurs by breaking the capsid open, and not by disassembly into individual subunits. Our data directly visualize a key step in HIV-1 replication and enhance our mechanistic understanding of the viral life cycle. - doi.org/10.1016/j.cell.2021.01.025 - 2021
S1 Ep 24PubReading [29] - Using antibodies to control DNA-templated chemical reactions - L. B. Pellejero, F. Ricci, T. Brown Jr et al.
DNA-templated synthesis takes advantage of the programmability of DNA-DNA interactions to accelerate chemical reactions under diluted conditions upon sequence-specific hybridization. While this strategy has proven advantageous for a variety of applications, including sensing and drug discovery, it has been so far limited to the use of nucleic acids as templating elements. Here, we report the rational design of DNA templated synthesis controlled by specific IgG antibodies. Our approach is based on the co-localization of reactants induced by the bivalent binding of a specific IgG antibody to two antigen-conjugated DNA templating strands that triggers a chemical reaction that would be otherwise too slow under diluted conditions. This strategy is versatile, orthogonal and adaptable to different IgG antibodies and can be employed to achieve the targeted synthesis of clinically-relevant molecules in the presence of specific IgG biomarker antibodies. - doi.org/10.1038/s41467-020-20024-3 - 2020
S1 Ep 23PubReading [28] - DNA G‐quadruplexes in the human genome: detection, functions and therapeutic potential - R. Hänsel-Hertsch, M. Di Antonio and S. Balasubramanian
Single-stranded guanine-rich DNA sequences can fold into four-stranded DNA structures called G-quadruplexes (G4s) that arise from the self-stacking of two or more guanine quartets. There has been considerable recent progress in the detection and mapping of G4 structures in the human genome and in biologically relevant contexts. These advancements, many of which align with predictions made previously in computational studies, provide important new insights into the functions of G4 structures in, for example, the regulation of transcription and genome stability, and uncover their potential relevance for cancer therapy. - doi:10.1038/nrm.2017.3 - 2017
S1 Ep 22PubReading [27] - Rapid Generation of Long Noncoding RNA Knockout Mice Using CRISPR/Cas9 Technology - N. R. Hansmeier, J.W. Kornfeld et al.
In recent years, long noncoding RNAs (lncRNAs) have emerged as multifaceted regulators of gene expression, controlling key developmental and disease pathogenesis processes. However, due to the paucity of lncRNA loss-of-function mouse models, key questions regarding the involvement of lncRNAs in organism homeostasis and (patho)-physiology remain difficult to address experimentally in vivo. The clustered regularly interspaced short palindromic repeats (CRISPR)/Cas9 platform provides a powerful genome-editing tool and has been successfully applied across model organisms to facilitate targeted genetic mutations, including Caenorhabditis elegans, Drosophila melanogaster, Danio rerio and Mus musculus. However, just a few lncRNA-deficient mouse lines have been created using CRISPR/Cas9-mediated genome engineering, presumably due to the need for lncRNA-specific gene targeting strategies considering the absence of open-reading frames in these loci. Here, we describe a step-wise procedure for the generation and validation of lncRNA loss-of-function mouse models using CRISPR/Cas9-mediated genome engineering. In a proof-of-principle approach, we generated mice deficient for the liver-enriched lncRNA Gm15441, which we found downregulated during development of metabolic disease and induced during the feeding/fasting transition. Further, we discuss guidelines for the selection of lncRNA targets and provide protocols for in vitro single guide RNA (sgRNA) validation, assessment of in vivo gene-targeting efficiency and knockout confirmation. The procedure from target selection to validation of lncRNA knockout mouse lines can be completed in 18–20 weeks, of which
S1 Ep 21PubReading [26] - Tools and methods for circular dichroism spectroscopy of proteins: a tutorial review - A. J. Miles, Robert W. Janes and B. A. Wallace
Circular dichroism (CD) spectroscopy is a widely-used method in biochemistry, structural biology and pharmaceutical chemistry. More than 24000 papers published in the past decade have included CD characterisations of proteins; many of those studies have also included other complementary chemical, biophysical, and computational chemistry methods. This tutorial review describes the background to the technique of CD spectroscopy and good practice methods for high quality data collection. It specifically focuses on both established and new methods and tools available for experimental design and interpretation, data processing, visualisation, analysis, validation, archiving, and accession, including tools developed to enhance the complementarity of this method with other structural and chemical biology studies. - DOI: 10.1039/d0cs00558d - 2021
S1 Ep 20PubReading [25] - Brown adipose tissue monocytes support tissue expansion - A. Gallerand, L. Yvan-Charvet1 & S. Ivanov et al.
Monocytes are part of the mononuclear phagocytic system. Monocytes play a central role during inflammatory conditions and a better understanding of their dynamics might open therapeutic opportunities. In the present study, we focused on the characterization and impact of monocytes on brown adipose tissue (BAT) functions during tissue remodeling. Single-cell RNA sequencing analysis of BAT immune cells uncovered a large diversity in monocyte and macrophage populations. Fate-mapping experiments demonstrated that the BAT macrophage pool requires constant replenishment from monocytes. Using a genetic model of BAT expansion, we found that brown fat monocyte numbers were selectively increased in this scenario. This observation was confirmed using a CCR2-binding radiotracer and positron emission tomography. Importantly, in line with their tissue recruitment, blood monocyte counts were decreased while bone marrow hematopoiesis was not affected. Monocyte depletion prevented brown adipose tissue expansion and altered its architecture. Podoplanin engagement is strictly required for BAT expansion. Together, these data redefine the diversity of immune cells in the BAT and emphasize the role of monocyte recruitment for tissue remodeling. - doi.org/10.1038/s41467-021-25616-1 - 2021
S1 Ep 19PubReading [24] - Root of the Tree: The Significance, Evolution, and Origins of the Ribosome - J. C. Bowman, L. D. Williams et al.
The ribosome is an ancient molecular fossil that provides a telescope to the origins of life. Made from RNA and protein, the ribosome translates mRNA to coded protein in all living systems. Universality, economy, centrality and antiquity are ingrained in translation. The translation machinery dominates the set of genes that are shared as orthologues across the tree of life. The lineage of the translation system defines the universal tree of life. The function of a ribosome is to build ribosomes; to accomplish this task, ribosomes make ribosomal proteins, polymerases, enzymes, and signaling proteins. Every coded protein ever produced by life on Earth has passed through the exit tunnel, which is the birth canal of biology. During the root phase of the tree of life, before the last common ancestor of life (LUCA), exit tunnel evolution is dominant and unremitting. Protein folding coevolved with evolution of the exit tunnel. The ribosome shows that protein folding initiated with intrinsic disorder, supported through a short, primitive exit tunnel. Folding progressed to thermodynamically stable β- structures and then to kinetically trapped α-structures. The latter were enabled by a long, mature exit tunnel that partially offset the general thermodynamic tendency of all polypeptides to form β-sheets. RNA chaperoned the evolution of protein folding from the very beginning. The universal common core of the ribosome, with a mass of nearly 2 million Daltons, was finalized by LUCA. The ribosome entered stasis after LUCA and remained in that state for billions of years. Bacterial ribosomes never left stasis. Archaeal ribosomes have remained near stasis, except for the superphylum Asgard, which has accreted rRNA post LUCA. Eukaryotic ribosomes in some lineages appear to be logarithmically accreting rRNA over the last billion years. Ribosomal expansion in Asgard and Eukarya has been incremental and iterative, without substantial remodeling of pre-existing basal structures. The ribosome preserves information on its history. - dx.doi.org/10.1021/acs.chemrev.9b00742 - 2020
S1 Ep 18PubReading [23] - Deformylation of 5-Formylcytidine in Different Cell Types - E. Korytiakovà and T. Carell
Epigenetic programming of cells requires methylation of deoxycytidines (dC) to 5-methyl-dC (mdC) followed by oxidation to 5-hydroxymethyl-dC (hmdC), 5-formyl-dC (fdC), and 5-carboxy-dC (cadC). Subsequent transformation of fdC and cadC back to dC by various pathways establishes a chemical intra-genetic control circle. One of the discussed pathways involves the Tdg-independent deformylation of fdC directly to dC. Here we report the synthesis of a fluorinated fdC feeding probe (F-fdC) to study direct deformylation to F-dC. The synthesis was performed along a novel pathway that circumvents any F-dC as a reaction intermediate to avoid contamination interference. Feeding of F-fdC and observation of F-dC formation in vivo allowed us to gain insights into the Tdg-independent removal process. While deformylation was shown to occur in stem cells, we here provide data that prove deformylation also in different somatic cell types. We also investigated active demethylation in a non-dividing neurogenin-inducible system of iPS cells that differentiate into bipolar neurons. - doi.org/10.1002/anie.202107089. - 2021
S1 Ep 17PubReading [22] - Shining Light on CRISPR Gene Editing - L. Taemaitree and T. Brown
Advanced spatiotemporal control of CRISPR-Cas9 activity is demonstrated through the use of chemically modified photoactivatable guide RNA. - doi.org/10.1021/acscentsci.0c00350 - 2020
S1 Ep 16PubReading [21] - DNA structure from A to B - R.Dickerson and HL. Ng
P. Shing Ho and his colleagues at Oregon State and Berkeley publish in this issue of PNAS an interesting study (1) of helical structure in the DNA hexamer GGCGCC, finding that various states that appear to be logical intermediates between A-DNA and B-DNA can be induced by methylation or bromination of cytosine or by crystal packing. Their results bear on three issues that have been argued over in the past: (i) the differences between A-DNA and B- DNA and transitions between them, (ii) the intrinsic sequence-dependent malleability of a DNA duplex, and (iii) the effects of local helix packing on DNA fine-structure. - doi-10.1073-pnas.141238898 - 2001
S2 Ep 5PubReading [20] - Anatomy, Back, Cauda Equina - E. Berg, J. Ashurst.
In 1595, French anatomist Andre du Laurens first described the structure of a rope-like tail of fibers at the caudal end of the spinal cord. This bundle of numerous axons was termed the cauda equina, from the Latin translation meaning “horse’s tail,” and it contains nerves which innervate both sensory and motor targets within lumbar, sacral, and coccygeal spinal cord levels. Epidemiologic assessments regard lesions to the cauda equina as uncommon, with a prevalence of 1 to 3 per 100,000 subjects. Typically caused by a herniated intervertebral disc at the L5-S1 levels, such lesions affect females as often as males and manifest as a number of urogenital and neuromuscular symptoms in the namesake “cauda equina syndrome." - StatPearls Publishing -2021
S2 Ep 4PubReading [19] - Hospitalization Length after Myocardial Infarction: Risk-Assessment-Based Time of Hospital Discharge vs. Real Life Practice - Michał Wegiel and Tomasz Rakowski
According to guidelines, it is safe for low-risk patients with myocardial infarction (MI) to be discharged within 72 h of hospitalization. However, results coming from registries show that the hospital stay is often much longer in a real-life situation. Data on the length of the hospital stay (LOS) of MI patients in Polish centers are lacking. We enrolled 212 consecutive patients with acute MI. Low-risk patients were defined according to PAMI II criteria: age 45%, no persistent ventricular arrhythmia, and no multi-vessel disease (MVD). The median of the hospitalization length was eight days (Q1: 6; Q3: 9). In low-risk patients (25%), the median of LOS was six days (Q1: 5; Q3: 7) (p regression analysis patients age, LVEF, ST-segment-elevation MI and the presence of MVD were independent predictors of longer hospitals stay (≥8 days). During follow up, there were no significant differences in the rates of clinical events between patients with shorter (
S1 Ep 15PubReading [18] - Controlling and enhancing CRISPR systems - H. Shivram, B. Cress, G. Knott, J. Doudna
Many bacterial and archaeal organisms use CRISPR-Cas (clustered regularly interspaced short palindromic repeats-CRISPR associated) systems to defend themselves from mobile genetic elements. These CRISPR-Cas systems are classified into six types based on their composition and mechanism. CRISPR-Cas enzymes are widely used for genome editing and offer immense therapeutic opportunity to treat genetic diseases. To realize their full potential, it is important to control the timing, duration, efficiency and specificity of CRISPR-Cas enzyme activities. In this review we discuss the mechanisms of natural CRISPR-Cas regulatory biomolecules that enhance or inhibit CRISPR-Cas immunity by altering enzyme function. We also discuss the potential applications of these CRISPR regulators and highlight unanswered questions about their evolution and purpose in nature. - doi:10.1038/s41589-020-00700-7 - 2021
S1 Ep 14PubReading [17] - Antisense, RNAi, and gene silencing strategies for therapy: Mission possible or impossible? - E. Rayburn and R. Zhang
Antisense oligonucleotides can regulate gene expression in living cells. As such, they regulate cell function and division, and can modulate cellular responses to internal and external stresses and stimuli. Although encouraging results from preclinical and clinical studies have been obtained and significant progress has been made in developing these agents as drugs, they are not yet recognized as effective therapeutics. Several major hurdles remain to be overcome, including problems with efficacy, off-target effects, delivery and side effects. The lessons learned from antisense drug development can help in the development of other oligonucleotide-based therapeutics such as CpG oligonucleotides, RNAi and miRNA. - doi:10.1016/j.drudis.2008.03.014 - 2008
S1 Ep 13PubReading [16] - Measuring cancer evolution from the genome - T. Graham and A. Sottoriva -
The temporal dynamics of cancer evolution remain elusive, because it is impractical to longitudinally observe cancers unperturbed by treatment. Consequently, our knowledge of how cancers grow largely derives from inferences made from a single point in time – the endpoint in the cancer’s evolution, when it is removed from the body and studied in the laboratory. Fortuitously however, the cancer genome, by virtue of ongoing mutations that uniquely mark clonal lineages within the tumour, provides a rich, yet surreptitious, record of cancer development. In this review, we describe how a cancer’s genome can be analysed to reveal the temporal history of mutation and selection, and discuss why both selective and neutral evolution feature prominently in carcinogenesis. We argue that selection in cancer can only be properly studied once we have some understanding of what the absence of selection looks like. We review the data describing punctuated evolution in cancer, and reason that punctuated phenotype evolution is consistent with both gradual and punctuated genome evolution. We conclude that, to map and predict evolutionary trajectories during carcinogenesis, it is critical to better understand the relationship between genotype change and phenotype change.- DOI: 10.1002/path.4821 - 2016
S1 Ep 12PubReading [15] - Engineering antibody therapeutics - M. Chiu and G. Gilliland
The successful introduction of antibody-based protein therapeutics into the arsenal of treatments for patients has within a few decades fostered intense innovation in the production and engineering of antibodies. Reviewed here are the methods currently used to produce antibodies along with how our knowledge of the structural and functional characterization of immunoglobulins has resulted in the engineering of antibodies to produce protein therapeutics with unique properties, both biological and biophysical, that are leading to novel therapeutic approaches. Antibody engineering includes the introduction of the antibody combining site (variable regions) into a host of architectures including bi and multi-specific formats that further impact the therapeutic properties leading to further advantages and successes in patient treatment. - http://dx.doi.org/10.1016/j.sbi.2016.07.012 - 2016
S2 Ep 3PubReading [14] - Acute abdominal pain in patients with Crohn’s disease: What urgent imaging tests should be done? - P. García, A. López-Jurado, A. Bártulos
Crohn’s disease is an autoimmune disease that predominantly affects the gastrointestinal tract. Crohn’s disease is diagnosed at a young age and runs a chronic course with acute flare-ups. When patients with Crohn’s disease present with flare-ups at the emergency department, they are usually managed in a way similar to patients with acute abdomen; there is no consensus about the most appropriate imaging work-up for patients with flare-ups of Crohn’s disease. Thus, we decided to review the literature about the imaging tests indicated (whether related to their diagnostic performance or to lower exposure to ionizing radiation) for acute flare-ups in patients with Crohn’s disease. - DOI: 10.1016/j.rx.2018.12.003 - 2019
S1 Ep 11PubReading [13] - Herpesvirus capsid assembly and DNA packaging - J. Heming, J. Conway and F. Homa
Herpes simplex virus type I (HSV-1) is the causative agent of several pathologies ranging in severity from the common cold sore to life-threatening encephalitic infection. During productive lytic infection, over 80 viral proteins are expressed in a highly regulated manner, resulting in the replication of viral genomes and assembly of progeny virions. The virion of all herpesviruses consists of an external membrane envelope, a proteinaceous layer called the tegument, and an icosahedral capsid containing the double-stranded linear DNA genome. The capsid shell of HSV-1 is built from four structural proteins: a major capsid protein, VP5, which forms the capsomers (hexons and pentons), the triplex consisting of VP19C and VP23 found between the capsomers, and VP26 which binds to VP5 on hexons but not pentons. In addition, the dodecameric pUL6 portal complex occupies one of the 12 capsid vertices, and the capsid vertex specific component (CVSC), a heterotrimer complex of pUL17, pUL25 and pUL36 binds specifically to the triplexes adjacent to each penton. The capsid is assembled in the nucleus where the viral genome is packaged into newly assembled closed capsid shells. Cleavage and packaging of replicated, concatemeric viral DNA requires the seven viral proteins encoded by the UL6, UL15, UL17, UL25, UL28, UL32, and UL33 genes. Considerable advances have been made in understanding the structure of the herpesvirus capsid and the function of several of the DNA packaging proteins by applying biochemical, genetic, and structural techniques. This review is a summary of recent advances with respect to the structure of the HSV-1 virion capsid and what is known about the function of the seven packaging proteins and their interactions with each other and with the capsid shell. - doi:10.1007/978-3-319-53168-7_6. - 2017
S1 Ep 10PubReading [12] - Ruthenium Complexes as Anticancer Agents: A Brief History and Perspectives - S. Lee, C. Kim, T.G. Nam
Platinum (Pt)-based anticancer drugs such as cisplatin have been used to treat various cancers. However, they have some limitations including poor selectivity and toxicity towards normal cells and increasing chemoresistance. Therefore, there is a need for novel metallo-anticancers, which has not been met for decades. Since the initial introduction of ruthenium (Ru) polypyridyl complex, a number of attempts at structural evolution have been conducted to improve efficacy. Among them, half-sandwich Ru-arene complexes have been the most prominent as an anticancer platform. Such complexes have clearly shown superior anticancer profiles such as increased selectivity toward cancer cells and ameliorating toxicity against normal cells compared to existing Pt-based anticancers. Currently, several Ru complexes are under human clinical trials. For improvement in selectivity and toxicity associated with chemotherapy, Ru complexes as photodynamic therapy (PDT), and photoactivated chemotherapy (PACT), which can selectively activate prodrug moieties in a specific region, have also been investigated. With all these studies on these interesting entities, new metallo- anticancer drugs to at least partially replace existing Pt-based anticancers are anticipated. This review covers a brief description of Ru-based anticancer complexes and perspectives. https://doi.org/10.2147/DDDT.S275007 - 2020
S2 Ep 2PubReading [11] - Outcomes of critically ill patients with acute kidney injury in COVID-19 infection: an observational study - Rodrigo Bezerra et al.
Background: Early reports indicate that AKI is common during COVID-19 infection. Different mortality rates of AKI due to SARS-CoV-2 have been reported, based on the degree of organic dysfunction and varying from public to private hospitals. However, there is a lack of data about AKI among critically ill patients with COVID-19.Methods: We conducted a multicenter cohort study of 424 critically ill adults with severe acute respiratory syndrome (SARS) and AKI, both associated with SARS-CoV-2, admitted to six public ICUs in Brazil. We used multivariable logistic regression to identify risk factors for AKI severity and in-hospital mortality.Results: The average age was 66.42±13.79years, 90.3% were on mechanical ventilation (MV), 76.6% were at KDIGO stage 3, and 79% underwent hemodialysis. The overall mortality was 90.1%. We found a higher frequency of dialysis (82.7% versus 45.2%), MV (95% versus 47.6%), vasopressors (81.2% versus 35.7%) (p
S1 Ep 9PubReading [9] - Principles of ubiquitin and SUMO modifications in DNA repair - S. Bergink, S. Jentsch
With the discovery in the late 1980s that the DNA-repair gene RAD6 encodes a ubiquitin-conjugating enzyme, it became clear that protein modification by ubiquitin conjugation has a much broader significance than had previously been assumed. Now, two decades later, ubiquitin and its cousin SUMO are implicated in a range of human diseases, including breast cancer and Fanconi anaemia, giving fresh momentum to studies focused on the relationships between ubiquitin, SUMO and DNA-repair pathways. - NATURE - 2009 -. doi:10.1038/nature07963
S2 Ep 1PubReading [10] - Recommendations on pre-hospital and early hospital management of acute heart failure - short version - Alexandre Mebazaa et al.
Despite several critical steps forward in the management of chronic heart failure (CHF), the area of acute heart failure (AHF) has remained relatively stagnant. As stated in the updated ESC HF guidelines, clinicians responsible for managing patients with AHF must frequently make treatment decisions without adequate evidence, usually on the basis of expert opinion consensus.2 Specifically, the treatment of acute HF remains largely opinion-based with little good evidence to guide therapy. Acute heart failure is a syndrome in which emergency physicians, cardiologists, intensivists, nurses, and other healthcare providers have to cooperate to provide ‘rapid’ benefit to the patients. We hereby would like to underscore the wider experience grown in different settings of the area of intensive care on acute heart failure, actually larger and more composite than that got in specialized Care Units. The distillate of such different experiences is discussed and integrated in the present document. Hence, the authors of this consensus paper believe a common working definition of AHF covering all dimensions and modes of presentations has to be made, with the understanding that most AHF presentations are either acute decompensations of chronic underlying HF or the abrupt onset of dyspnoea associated with significantly elevated blood pressure. Secondly, recent data show that, much like acute coronary syndrome, AHF might have a ‘time to therapy’ concept. Accordingly, ‘pre-hospital’ management is considered a critical component of care. Thirdly, most patients with AHF have normal or high blood pressure at presentation and are admitted with symptoms and/or signs of congestion. This is in contradiction to the presentation where low cardiac output leads to symptomatic hypotension and signs/symptoms of hypoperfusion, a circumstance that is relatively rare, present in coronary care unit/ intensive care unit (CCU/ICU) but associated with a particularly poor outcome. Hence, it is important to note that appropriate therapy requires appropriate identification of the specific AHF pheno-type.3 The aim of the current paper is not to replace guidelines, but, to provide contemporary perspective for early hospital management within the context of the most recent data and to provide guidance, based on expert opinions, to practicing physicians and other healthcare professionals (Figure 1). We believe that the experience accrued in the different settings from the emergency department through to the ICU/CCU is collectivel valuable in determining how best to manage the patients with AHF. Herein, a shortened version mainly including group recommendations is provided. Full version of the consensus paper is provided as Supplementary material online. - Alexandre Mebazaa - http://onlinelibrary.wiley.com/doi/10.1002/ejhf.289/full - 2015
S1 Ep 8PubReading [8] - A Glimpse of Structural Biology through X-Ray Crystallography - Yigong Shi
Since determination of the myoglobin structure in 1957, X-ray crystallography, as the anchoring tool of structural biology, has played an instrumental role in deciphering the secrets of life. Knowledge gained through X-ray crystallography has fundamentally advanced our views on cellular processes and greatly facilitated development of modern medicine. In this brief narrative, I describe my personal understanding of the evolution of structural biology through X-ray crystallography—using as examples mechanistic understanding of protein kinases and integral membrane proteins—and comment on the impact of technological development and outlook of X-ray crystallography - dx.doi.org/10.1016/j.cell.2014.10.051 - 2014
S1 Ep 7PubReading [7] - Modified nucleic acids: replication, evolution, and next-generation therapeutics - K. Duffy, S. Arangundy-Franklin and P. Holliger
Modified nucleic acids, also called xeno nucleic acids (XNAs), offer a variety of advantages for biotechnological applications and address some of the limitations of first-generation nucleic acid therapeutics. Indeed, several therapeutics based on modified nucleic acids have recently been approved and many more are under clinical evaluation. XNAs can provide increased biostability and furthermore are now increasingly amenable to in vitro evolution, accelerating lead discovery. Here, we review the most recent discoveries in this dynamic field with a focus on progress in the enzymatic replication and functional exploration of XNAs. K. Duffy, S. Arangundy-Franklin and P. Holliger -doi.org/10.1186/s12915-020-00803-6 - 2020
S1 Ep 6PubReading [6] Will CRISPR‐Cas9 Have Cards to Play Against Cancer? An Update on its Applications - P Daisy, K Shreyas, T. S. Anitha - Molecular Biotechnology
Genome editing employs targeted nucleases as powerful tools to precisely alter the genome of target cells and regulate functional genes. Various strategies have been risen so far as the molecular scissors-mediated genome editing that includes zinc finger nuclease, transcription activator-like effector nucleases, and clustered regularly interspaced short palindromic repeats—CRISPR-related protein 9. These tools allow researchers to understand the basics of manipulating the genome, create animal models to study human diseases, understand host–pathogen interactions and design disease targets. Targeted genome modification utilizing RNA-guided nucleases are of recent curiosity, as it is a fast and effective strategy that enables the researchers to manipulate the gene of interest, carry out functional studies, understand the molecular basis of the disease and design targeted therapies. CRISPR-Cas9, a bacterial defense system employed against viruses, consists of a single-strand RNA-guided Cas9 nuclease connected to the corresponding complementary target sequence. This powerful and versatile tool has gained tremendous attention among the researchers, owing to its ability to correct genetic disorders. To help illus- trate the potential of this gene editor in unexplored corners of oncology, we describe the history of CRISPR-Cas9, its rapid progression in cancer research as well as future perspectives. - P Daisy, K Shreyas, T. S. Anitha -doi.org/10.1007/s12033-020-00289-1 - 2021
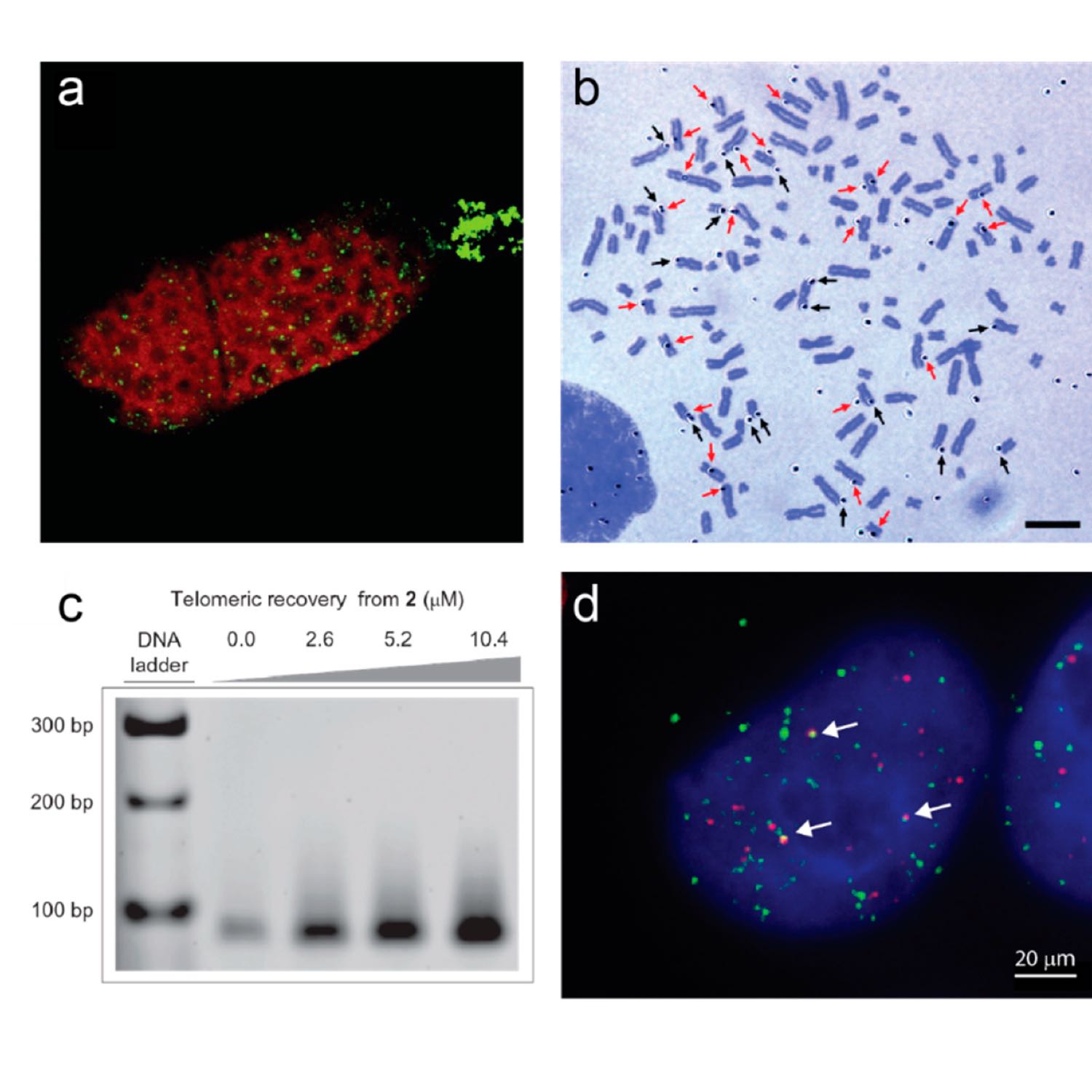
S1 Ep 5PubReading [5] G-Quadruplexes at Telomeres: Friend or Foe? - Tracy Bryan - Molecules
Telomeres are DNA-protein complexes that cap and protect the ends of linear chromosomes. In almost all species, telomeric DNA has a G/C strand bias, and the short tandem repeats of the G-rich strand have the capacity to form into secondary structures in vitro, such as four-stranded G-quadruplexes. This has long prompted speculation that G-quadruplexes play a positive role in telomere biology, resulting in selection for G-rich tandem telomere repeats during evolution. There is some evidence that G-quadruplexes at telomeres may play a protective capping role, at least in yeast, and that they may positively affect telomere maintenance by either the enzyme telomerase or by recombination-based mechanisms. On the other hand, G-quadruplex formation in telomeric DNA, as elsewhere in the genome, can form an impediment to DNA replication and a source of genome instability. This review summarizes recent evidence for the in vivo existence of G-quadruplexes at telomeres, with a focus on human telomeres, and highlights some of the many unanswered questions regarding the location, form, and functions of these structures. - Tracy Bryan - doi:10.3390/molecules25163686 - 2020
S1 Ep 4PubReading [4] Triplex-forming oligonucleotides: a third strand for DNA nanotechnology - Arun Chandrasekaran, David Rusling - Nucleic Acids Research
DNA self-assembly has proved to be a useful bottom- up strategy for the construction of user-defined nanoscale objects, lattices and devices. The design of these structures has largely relied on exploiting simple base-pairing rules and the formation of double-helical domains as secondary structural elements. However, other helical forms involving specific non-canonical base-base interactions have introduced a novel paradigm into the process of engineering with DNA. The most notable of these is a three-stranded complex generated by the binding of a third strand within the duplex major groove, generating a triple-helical (‘triplex’) structure. The sequence, structural and assembly requirements that differentiate triplexes from their duplex counterparts has allowed the design of nanostructures for both dynamic and/or structural purposes, as well as a means to target non-nucleic acid components to precise locations within a nanostructure scaffold. Here, we review the properties of triplexes that have proved useful in the engineering of DNA nanostructures, with an emphasis on applications that hitherto have not been possible by duplex formation alone. Arun Chandrasekaran, David Rusling - doi: 10.1093/nar/gkx1230 - 2018
S1 Ep 3PubReading [3] DNA Structure: A-, B- and Z-DNA Helix Families - David W Ussery - Encyclopedia Of Life Science
There are three major families of DNA helices: A-DNA, B-DNA and Z-DNA. The helicalstructure of DNA is variable and depends on the sequence as well as the environment. David W Ussery
S1 Ep 2PubReading [2] New Approaches Towards Recognition of Nucleic Acid Triple Helices - Dev P. Arya - Acc Chem Res
We show that groove recognition of nucleic acid triple helices can be achieved with aminosugars. Among these aminosugars, neomycin is the most effective aminoglycoside (groove binder) for stabilizing a DNA triple helix. It stabilizes both the T·A·T triplex and mixed-base DNA triplexes better than known DNA minor groove binders (which usually destabilize the triplex) and polyamines. Neomycin selectively stabilizes the triplex (T·A·T and mixed base) without any effect on the DNA duplex. The selectivity of neomycin likely originates from its potential and shape complementarity to the triplex Watson–Hoogsteen groove, making it the first molecule that selectively recognizes a triplex groove over a duplex groove. The groove recognition of aminoglycosides is not limited to DNA triplexes, but also extends to RNA and hybrid triple helical structures. Intercalator–neomycin conjugates are shown to simultaneously probe the base stacking and groove surface in the DNA triplex. Calorimetric and spectrosocopic studies allow the quantification of the effect of surface area of the intercalating moiety on binding to the triplex. These studies outline a novel approach to the recognition of DNA triplexes that incorporates the use of non-competing binding sites. These principles of dual recognition should be applicable to the design of ligands that can bind any given nucleic acid target with nanomolar affinities and with high selectivity. Dev P. Arya

S1 Ep 1PubReading [1] A structure for Deoxyribose Nucleic Acid - J. D. Watson, F. H. C. Crick - Nature
Molecular Structure of Nucleic Acids - A structure for Deoxyribose Nucleic Acid - We wish to suggest astructure for the salt of deoxyribose nucleic acid (D.N.A.). This structure has novel features which are of considerable biological interest. J. D. Watson, F. H. C Crick.